DMD Gene Ontology

The AmiGO gene ontology database was used to find GO terms associated with the human DMD gene product (dystrophin). A list of the dystrophin-associated GO terms organized by ontology is shown below. The original list, along with the ability to graphically view each individual term, can be found by clicking this link.
GO TERM Accession Ontology
muscle homeostasis GO:0046716 biological process
muscle organ development GO:0007517 biological process
neuron projection morphogenesis GO:0048812 biological process
neurotransmitter receptor metabolic process GO:0045213 biological process
peptide biosynthetic process GO:0043043 biological process
regulation of transcription GO:0045449 biological process
skeletal muscle tissue development GO:0007519 biological process
cell surface GO:0009986 cellular component
cell-substrate junction GO:0030055 cellular component
costamere GO:0043034 cellular component
cytoplasm GO:005737 cellular component
cytoskeleton GO:0005856 cellular component
dystrophin-associated glycoprotein complex GO:0016010 cellular component
insoluble fraction GO:0005626 cellular component
membrane raft GO:0045121 cellular component
microsome GO:0005792 cellular component
nucleus GO:0005634 cellular component
sarcolemma GO:0042383 cellular component
synapse GO:0045202 cellular component
Z disc GO:0030018 cellular component
actin binding GO:0003779 molecular function
calcium ion binding GO:0005509 molecular function
nitric-oxide synthase binding GO:0050998 molecular function
PDZ domain binding GO:0030165 molecular function
protein binding GO:0005515 molecular function
structural constituent of cytoskeleton GO:0005200 molecular function
structural constituent of muscle GO:0008307 molecular function
zinc ion binding GO:0008270 molecular function
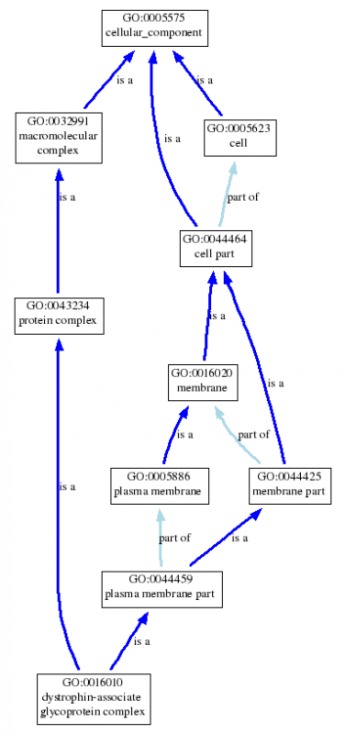
Below, is a graphical representation of the dystrophin-associated glycoprotein complex term. The complex is defined as "a multiprotein complex that forms a strong mechanical link between the cytoskeleton and extracellular matrix; typical of, but not confined to, muscle cells. The complex is composed of transmembrrane, cytoplasmic, and extracellular proteins, including dystrophin, sarcoglycans, dystroglycan, dystrobrevins, syntrophins, sarcospan, caveolin-3, and NO synthase". It, also, was obtained from AmiGO. Clicking on the picture will bring up the tree in its full size.
GO TERM Accession Ontology
muscle homeostasis GO:0046716 biological process
muscle organ development GO:0007517 biological process
neuron projection morphogenesis GO:0048812 biological process
neurotransmitter receptor metabolic process GO:0045213 biological process
peptide biosynthetic process GO:0043043 biological process
regulation of transcription GO:0045449 biological process
skeletal muscle tissue development GO:0007519 biological process
cell surface GO:0009986 cellular component
cell-substrate junction GO:0030055 cellular component
costamere GO:0043034 cellular component
cytoplasm GO:005737 cellular component
cytoskeleton GO:0005856 cellular component
dystrophin-associated glycoprotein complex GO:0016010 cellular component
insoluble fraction GO:0005626 cellular component
membrane raft GO:0045121 cellular component
microsome GO:0005792 cellular component
nucleus GO:0005634 cellular component
sarcolemma GO:0042383 cellular component
synapse GO:0045202 cellular component
Z disc GO:0030018 cellular component
actin binding GO:0003779 molecular function
calcium ion binding GO:0005509 molecular function
nitric-oxide synthase binding GO:0050998 molecular function
PDZ domain binding GO:0030165 molecular function
protein binding GO:0005515 molecular function
structural constituent of cytoskeleton GO:0005200 molecular function
structural constituent of muscle GO:0008307 molecular function
zinc ion binding GO:0008270 molecular function
Below, is a graphical representation of the dystrophin-associated glycoprotein complex term. The complex is defined as "a multiprotein complex that forms a strong mechanical link between the cytoskeleton and extracellular matrix; typical of, but not confined to, muscle cells. The complex is composed of transmembrrane, cytoplasmic, and extracellular proteins, including dystrophin, sarcoglycans, dystroglycan, dystrobrevins, syntrophins, sarcospan, caveolin-3, and NO synthase". It, also, was obtained from AmiGO. Clicking on the picture will bring up the tree in its full size.
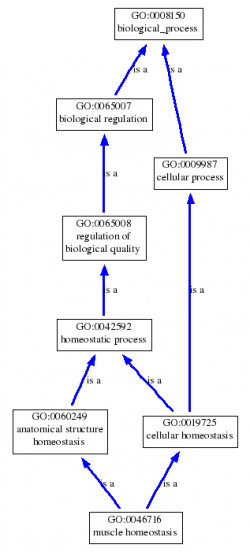
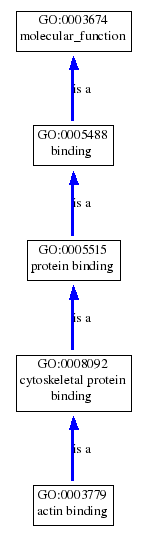
Below, is a graphical representation of the muscle homeostasis and actin binding GO terms for dystrophin. These GO terms illustrate that dystrophin is required for muscle structure maintenance specifically through its ability to bind actin (and connect it to the extra cellular matrix in muscle fibers). Clicking on the pictures will bring up the trees in their full sizes.
References
Carbon S, Ireland A, Mungall CJ, Shu S, Marshall B, Lewis S, AmiGO Hub, Web Presence Working Group. AmiGO: online access to ontology and annotation data. Bioinformatics. Jan 2009;25(2):288-9.
This site was created by Anthony Parendo for an undergraduate Genetics 677 course at the University of Wisconsin-Madison.